Handbook |
Search Handbook
|
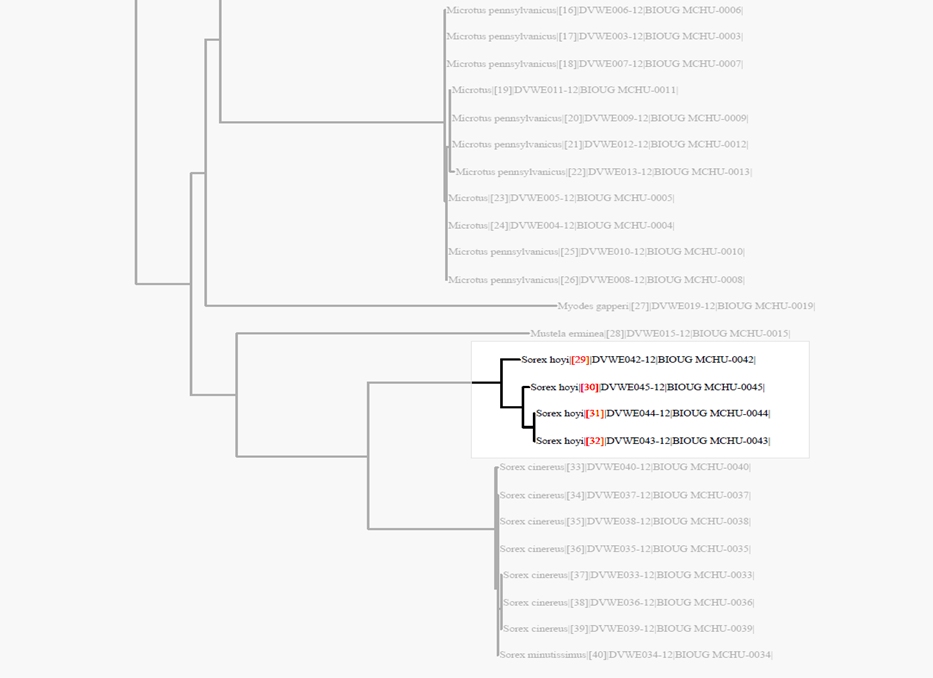
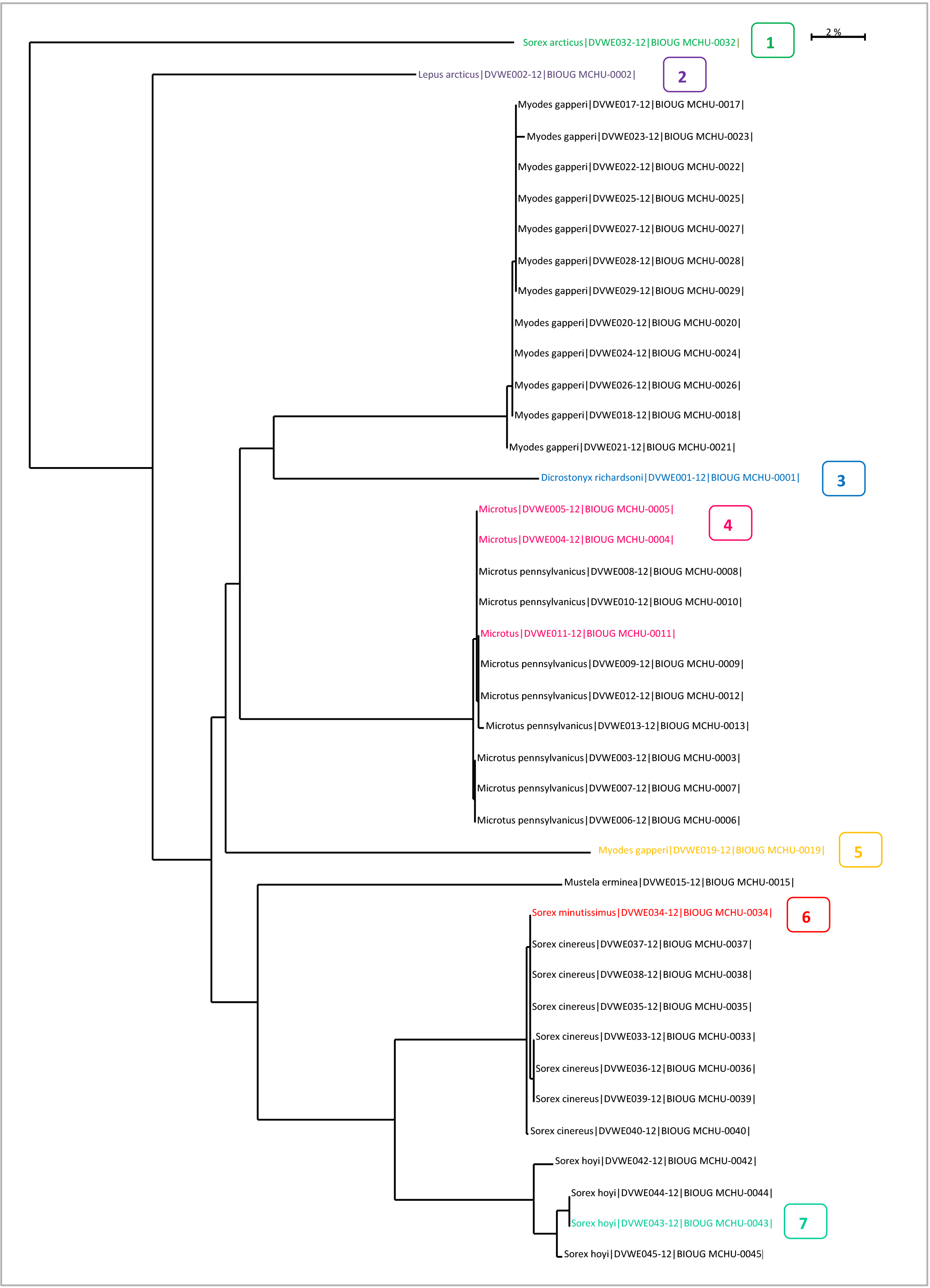
Taxon ID Trees for ValidationThe Taxon ID Tree on BOLD is a useful tool to identify problem sequences. Seven cases are described below.
Case 1: False outgroup resulted from a contaminationAn outgroup may be caused by contamination or it may be a real phenomenon resulting from a genetically distant taxa. The only way to know if an outgroup is the result of a contaminant is by comparing the nucleotide sequence to the BOLD ID engine database. To run a sequence against the BOLD ID engine:
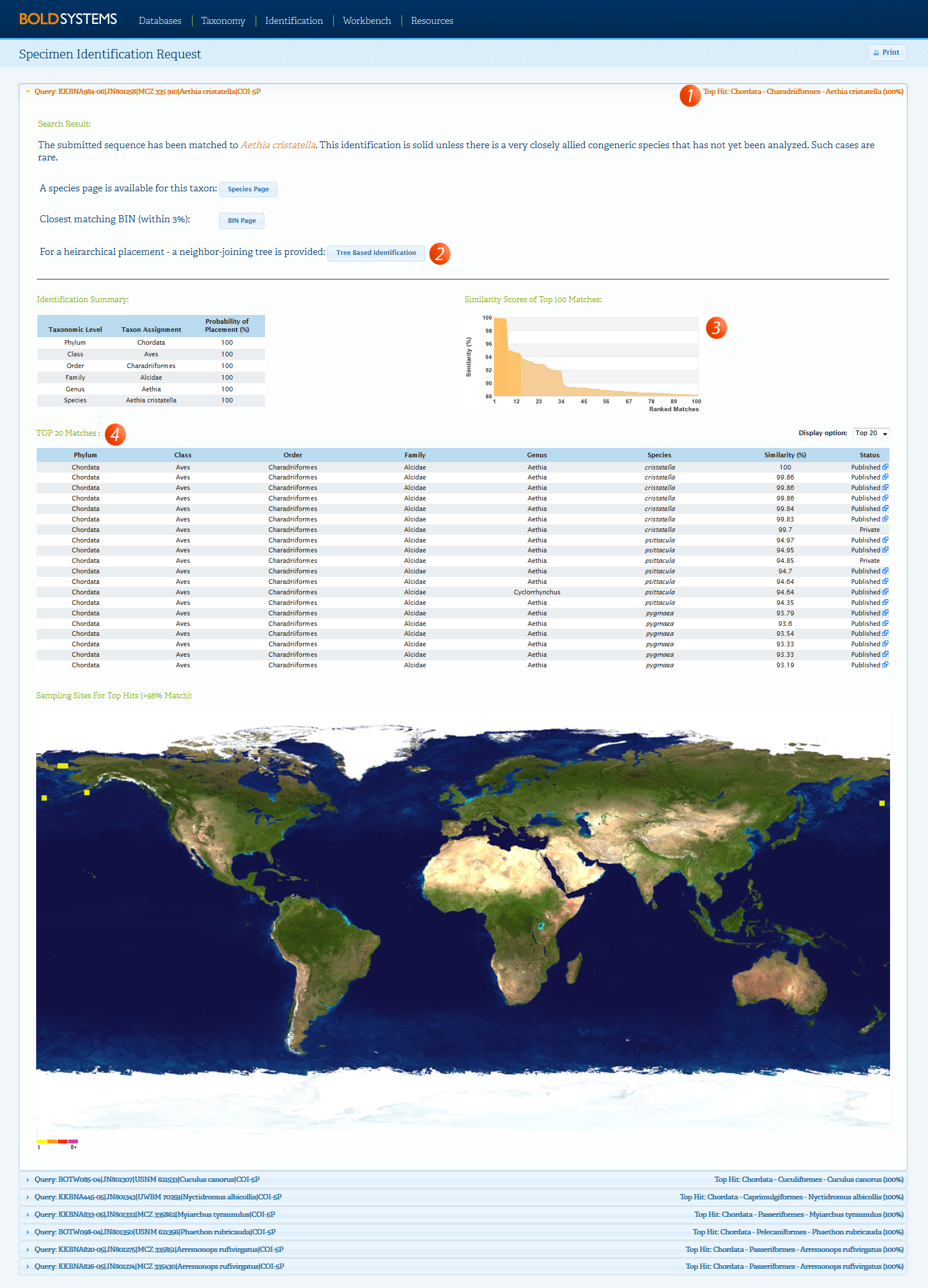
The Specimen Identification Request window will appear illustrating the top similarity matches as illustrated below. When the top match is at 99% similarity or higher and it does not agree with the taxonomic name provided, it usually indicates a contamination.
In this case, users can add annotation to the record to indicate the possible presence of a contamination. To add annotation to the affected records:
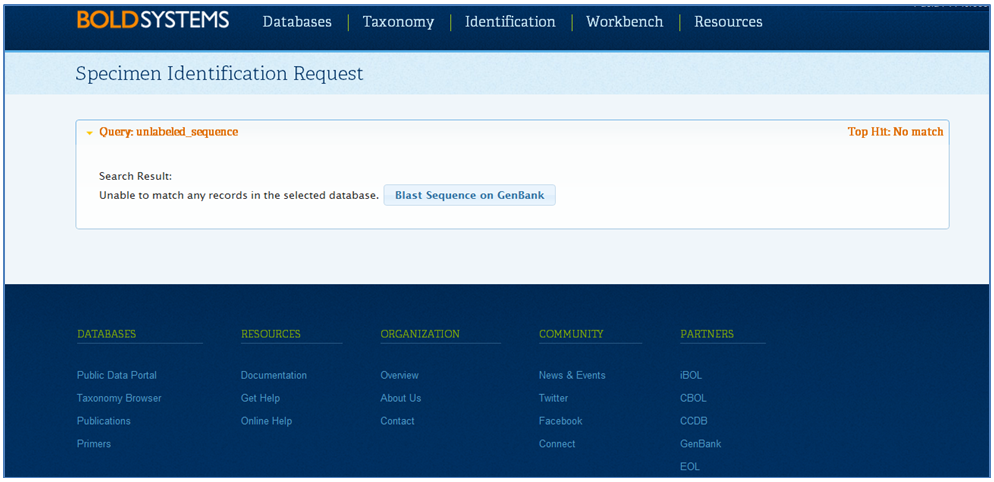
Case 2: Real outgroup resulted from a genetically unrelated taxonReal outgroups can sometimes be included on a tree. In order to determine if an outgroup is real or a contaminant, the sequence needs to be blasted against the BOLD ID Engine (refer to Case 1 - False outgroup resulted from a contamination for instructions on how to access the Identification Engine). If the outgroup represents a species new to BOLD, no record match will be displayed. The records should then be blasted against GenBank, which can be done directly from BOLD. When the BOLD ID Engine fails to find a match, click Blast Sequence on GenBank to directly access the Standard Nucleotide BLAST on GenBank. If the resulting identification on GenBank matches the name provided in the tree by more than 99%, it can be concluded that the identification is correct. This is a real outgroup and does not need to be tagged.
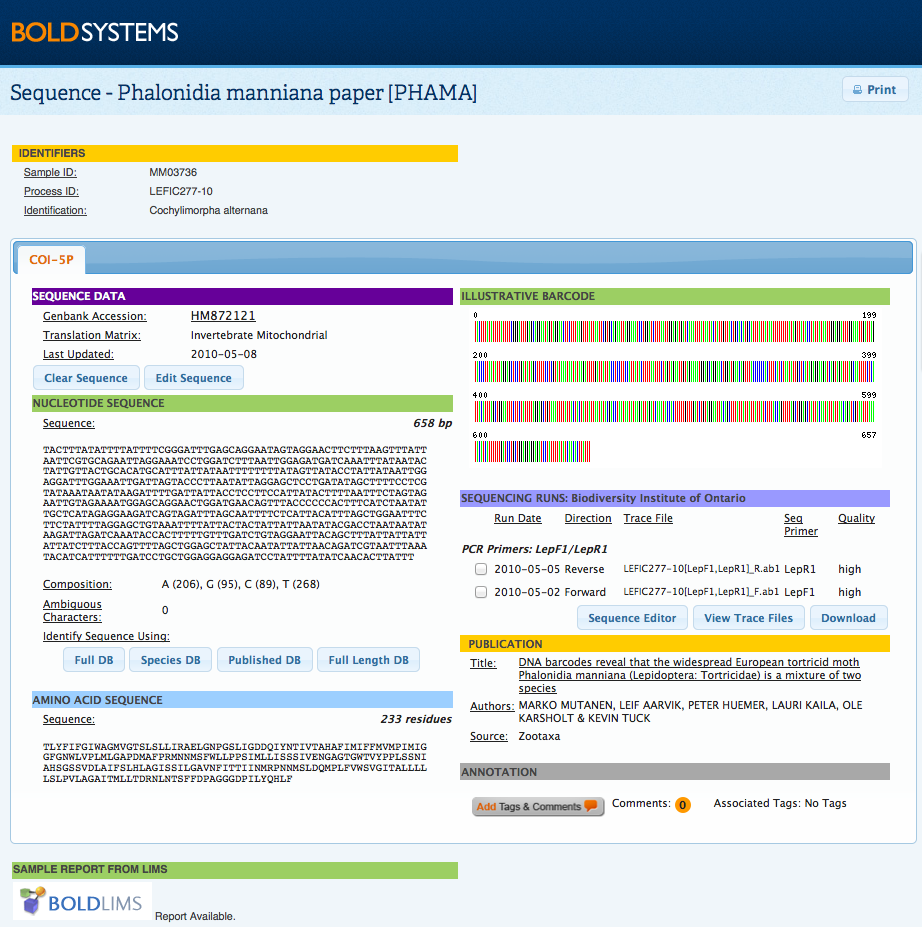
Case 3: Single branch resulted from unique recordSome species or haplotypes may appear as a single branch on the tree. It is important to check the identification of all single branches in a tree since these cannot be compared with other records within the same cluster. The Barcode Index Numbers (BIN) database can be used to confirm an identification; if the sequence meets the requirements to be clustered into BINs, then the record will have a BIN number. Refer to the Barcode Index Numbers (BIN) section in the Handbook. To navigate to the BIN page:
The data provided on the BIN page may help confirm the identity of single branch records on the tree, if other members of that species appear in other projects on BOLD. Where the correct identity of a single branch records cannot be confirmed right away, it is suggested that users monitor the BIN page for records over time as new specimens are being added to BOLD continously and activity on a BIN page is fluid.
Case 4: Incomplete identification on a clusterSome clusters on the tree may contain records that are identified to species and records that are not. It is possible to add full taxonomy to these records based on the tree and BOLD ID engine by sending a taxonomy update through the BOLD Submission Protocol. Tips and TroubleshootingWhen updating the taxonomy of a record based on the results from the identification engine, the Identified By field should be updated to "BOLD ID Engine". This informs other users that the identification provided was based on the record's nucleotide sequence without further examination of the voucher specimen and it should be reviewed by a taxonomic expert when possible. Further notes about taxonomic identifications can be added to the Taxonomy Notes and Identification Method fields. Case 5: Single branch resulted from contamination or misidentificationWhen two or more records with the same species name appear on a tree in separate branches, it is often the result of a contamination or misidentification. If a misidentification can be concluded and the correct identification is known, it is recommended that the taxonomy be updated as soon as possible without tagging the record. If a misidentification is not certain or the correct name is unknown, the record should be tagged and re-examined in the future. How to access record annotation:
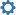
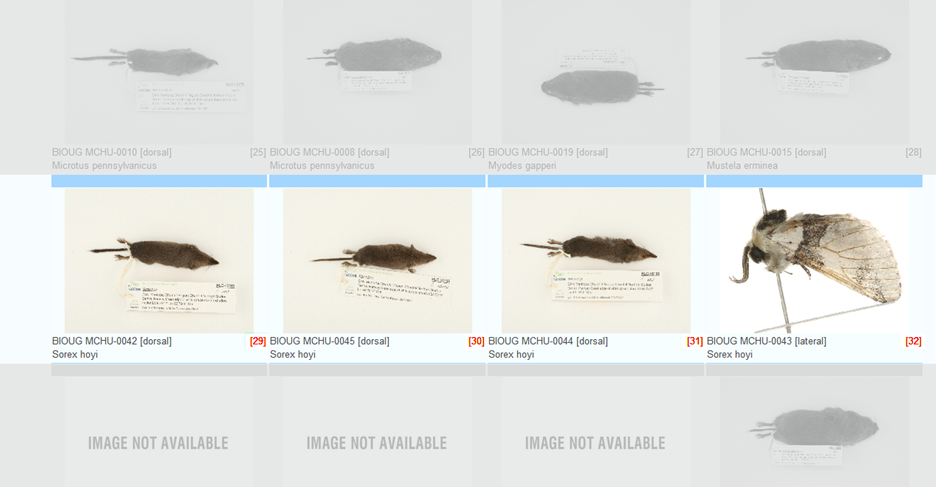
Case 6: Misidentified record in a cluster Some species can be difficult to identify solely on morphological characteristics. Sometimes Taxon ID trees can cluster together records that were believed to belong to two or more species. In certain cases this can be easily resolved by updating the taxonomy of misidentified records. Refer to the section on Updating Specimen Data. Tips and TroubleshootingBefore updating the taxonomy of any record in a project is important to check the sequence against the Identification Engine or BIN records (refer to the section on Identification Engine and BINs in this handbook) to ensure that the correct nomenclature matches other records on BOLD. Case 7: Image mismatchA mismatched image occurs when an incorrect picture is associated with a record. It is recommended to always create a matching image library when building a tree to examine records for this possible issue. Refer to the section on Taxon ID Trees in this handbook for more instructions on how to build a tree with matching images. When building the tree, choose "Matching Images and Spreadsheet" in the parameters window. Then from the Tree Result window choose the option to View Image List. Each branch on the tree will be automatically assigned a number that will correspond to a photo in the image library. See the screenshots below for an illustration.
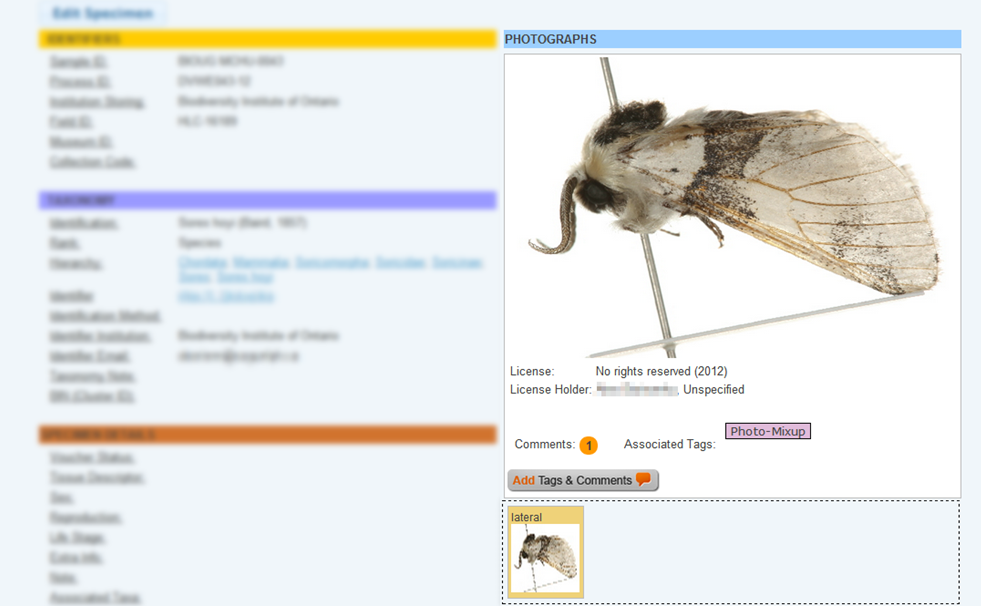
To correct an image mismatch:
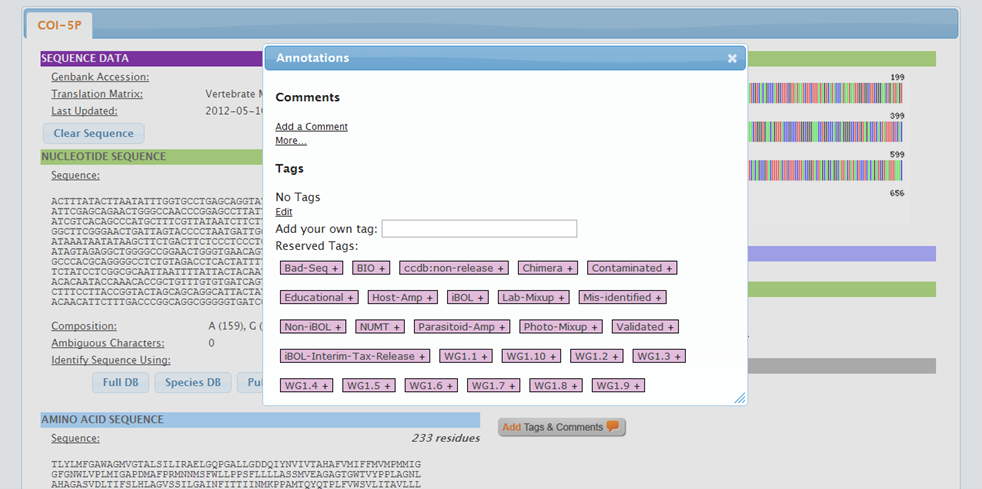
If the image mismatch cannot be resolved immediately, add a tag to the image to inform other users that this issue has been acknowledged. To add a tag on an image:
Back to Top |